Vanderbilt Researcher Identifies Modes of Natural Selection in Understudied Human Populations
By Andy Flick, Evolutionary Studies scientific coordinator
Amanda Lea, assistant professor of biology, along with a global team of experts, have discovered new pathways of natural selection in humans. The group, led by Tsimane Health and Life History Project co-Director Michael Gurven of the University of California, Santa Barbara, studied two populations of Bolivians that live in the lowlands northeast of the Andes mountains, the Tsimane and the Moseten. Previous work has shown that these tropical populations are exposed to high burdens of pathogens and parasites, while showing minimal cardiovascular disease or dementia. Results from this study suggest that the Tsimane population’s genome has undergone selection on traits associated with both immunity and metabolism, and that these selected regions are associated with health in the population today.
This research builds context into the evolution and specialization of humans across the globe. A few well-known examples of evolved human-specialization already exist. For example, some European, African, and Middle Eastern descendants can digest lactase which allows for milk drinking well past puberty in those populations.
According to Lea, “despite great interest in understanding the DNA variants that differ between populations in beneficial or adaptive ways, they remain very hard to identify and characterize. Nevertheless, doing so has the potential to uncover previously undescribed loci involved in evolutionarily- and biomedically-relevant traits.”
One especially unique piece of this project is that the study stems from a long-term partnership between the Tsimane Health and Life History Project and the Tsimane and Moseten communities. Over the past 20 years, the project has included collaborations between anthropologists, physicians, indigenous leadership, community members and local institutions to both build a comprehensive picture of health and to improve it. Such trusting collaborations are vital for navigating the ethical landscape surrounding the study of indigenous DNA.
On the importance of working with Bolivian Amerindians, Gurven said, “most studies so far that have linked DNA mutations to health have focused on Europeans or Americans. Groups like the Tsimane have a distinct history living in a distinct environment. It is likely that there are genetic signatures in the Tsimane genome that do not exist in most populations typically studied. Exploring novel genes and their relation to a wide range of health outcomes is something we are currently doing and are very excited about.”
Another, somewhat more surprising result from this research was that there appear to be beneficial genetic variants that affect the way Tsimane use energy. This is exciting, in light of the team’s prior work that showed that harmful alleles in the U.S. and Europe, like Apolipoprotein-E4 elevating risk of Alzheimer’s Disease, had surprising benefits in the high-infection and energy-limited context of Tsimane ecology. Further study of how genes affect immunity, infection risk and cardiometabolic health among Tsimane can shed new light on why certain deleterious alleles are still with us today. Lea explained that the Tsimane have exceptionally healthy hearts and a future project from this research will be to identify if the genetic differences the team found in energy usage relate to their cardiovascular and cerebral health.
This project was completed by working with the Tsimane, one of the many (>35) indigenous groups of Bolivia. The population of the Tsimane is around 17,000 people spread across roughly 90 villages. Blood samples were collected from more than 1,000 individuals between 2006 and 2015 by the Tsimane Health and Life History Project. DNA and SNP data were collected from the blood samples and the team identified genomic regions under positive selection. They then used statistical software and online databases to identify correlations in genetic regions known to be related to immunity. The group also tested the relationships between genotypes (genes) and phenotypes (how those genes are expressed).
Among other future directions, the team is poised to exploreother ways that Tsimane genetics are variable compared to that of other groups, such as Europeans. They are also looking to build a comprehensive health tapestry for the Tsimane using other biomedical research methods — transcriptomics, metabolomics, and microbiome — linking genetics to health.
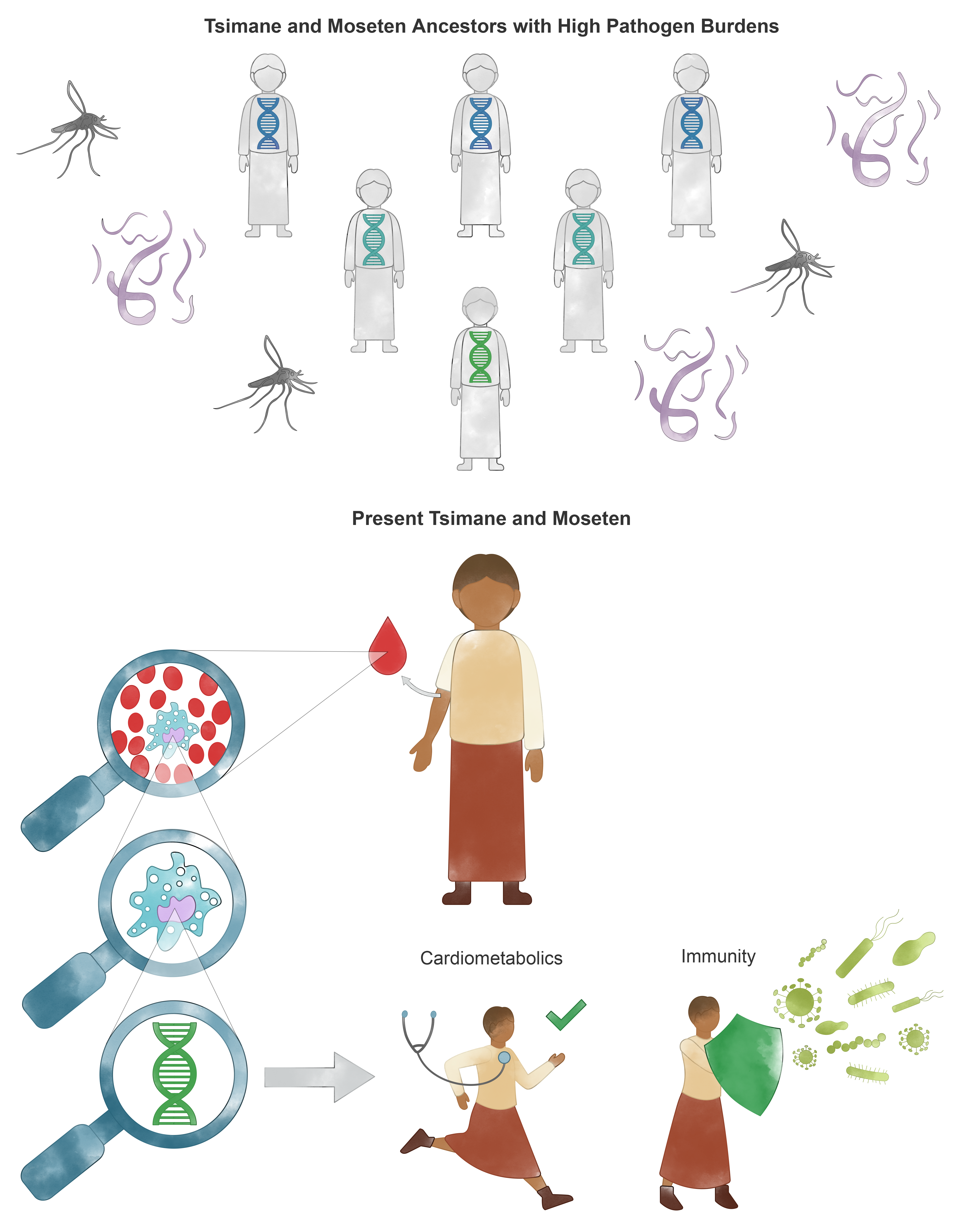
Top panel: The Tsimane and Moseten populations have experienced a diverse pathogen burden over many generations. Our findings suggest that this burden has selected for genetic variants involved in immune defense, and metabolism. Bottom panel: Regions that have been under selection over evolutionary time are associated with variation in health-related traits in the Tsimane today (specifically, biomarkers of immune and cardiometabolic health). Credit: This image was created by Caitlyn Skelton at Designs That Cell, with input from Amanda Lea and Michael Gurven. It is being used to communicate the results of our study with Tsimane and Moseten communities.
Funding: This work was supported by the National Institutes of Health grants R01AG024119, R56AG024119, and RF1AG054442. This project was made possible by the Tsimane Health and Life History Project and the Tsimane and Moseten communities.
Citation: Lea, A.J., Garcia, A., Arevalo, J., Ayroles, J.F., Buetow, K., Cole, S.W., Eid Rodriguez, D., Gutierrez, M., Highland, H.M., Hooper, P.L. and Justice, A., 2023. Natural selection of immune and metabolic genes associated with health in two lowland Bolivian populations. Proceedings of the National Academy of Sciences, 120(1), p.e2207544120.