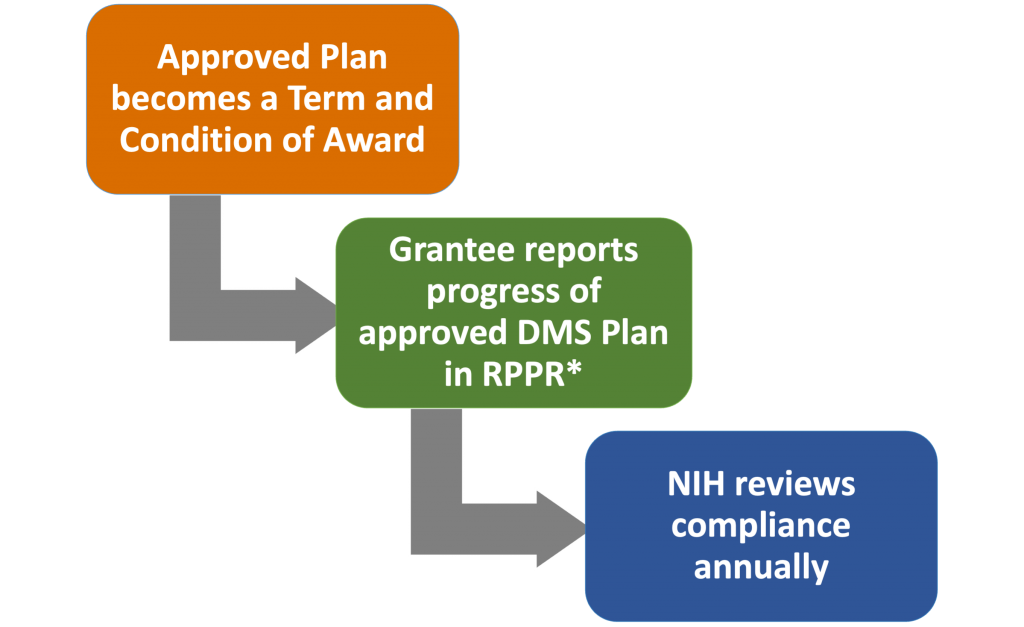
All National Institutes of Health (NIH) research grant proposals, both new and competing renewals, need to include a Data Management and Sharing plan, have costs accounted for in the budget, and have progress shared with NIH in annual reports. This page describes the Data Management and Sharing policy and makes recommendations regarding how our faculty can implement it for their NIH projects. You can send questions to ric@vanderbilt.edu.
Policy and Guidance
The new policy and guidance was presented to schools, centers, and staff throughout the winter of 2023. To request a presentation for your department or research group, please contact ric@vanderbilt.edu.
Overview
This policy applies only to research grants, not training grants, fellowships, infrastructure grants, instrument grants, nor non-competitive renewals.
The policy dictates how data generated using support from these grants must be managed and shared. Scientific Data is defined as “recorded factual material commonly accepted in the scientific community as of sufficient quality to validate and replicate research findings, regardless of whether the data are used to support scholarly publications.” The NIH definition excludes “data not necessary (or of sufficient quality) to validate and replicate research findings,” laboratory notebooks, preliminary analyses, and physical objects.
All new and competing grant proposals must include a plan based on the NIH DMS form template.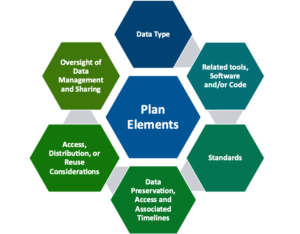
The plan must cover six elements:
- Data Type
- Related tools, software, and/or code
- Standards
- Data Preservation, access, and associated timelines
- Access, distribution, and reuse considerations
- Oversight of Data Management & Sharing
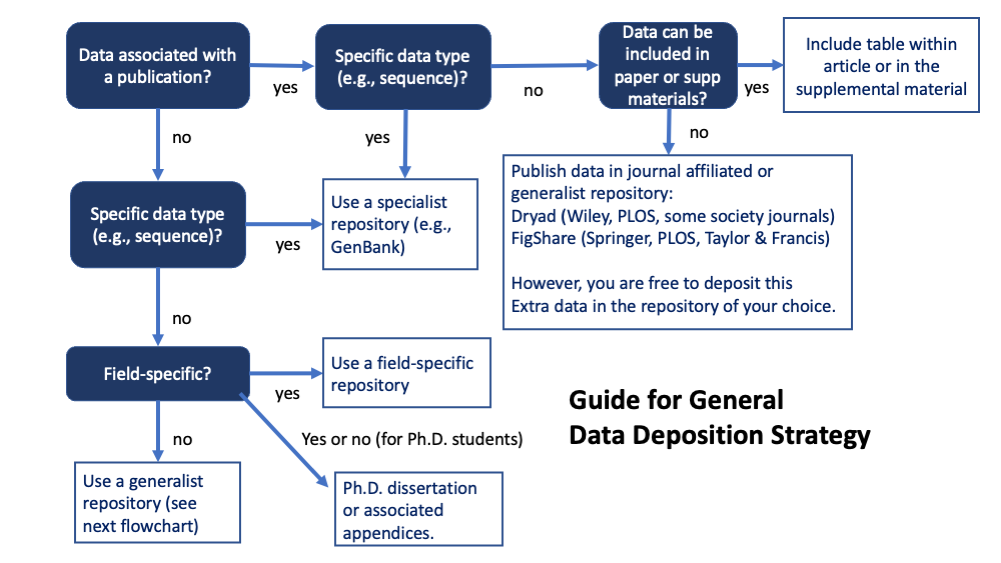
PIs are expected to maximize sharing, which typically mean depositing data and associated metadata into publicly accessible repositories, which in all cases must be associated with a “persistent unique identifier,” usually a web-based DOI code. The goal is that data should be deposited in a way that complies with the “F.A.I.R.” principle (findable, accessible, interoperable, and reusable).
Data must be shared at time of first publication or at the end of the project period, whichever comes first. Unpublished data (that meet the definition above) must be deposited and reported at end of project period even if they will end up in a published paper.
- Some grant programs, Institutes, Offices, or FOA may indicate particular data repositories to be used – follow any special instructions.
- Prioritize using established discipline or data-type specific repositories to make it easy for people in your field to find (PDB, Genbank, etc.) This may include data that is part of a publication or it’s supporting/supplementary materials, as well as online pre-prints (e.g., BioRxiv). NIH supports many Scientific Data Repositories and you can also search the Registry of Research Data Repositories.
- For small data sets generated by a graduate student, you may be able to include data in their Ph.D. dissertation (including appendix), which eventually will be assigned a DOI.
- Otherwise, use “generalist” repositories.
- Not all repositories are the same, so make sure you select on that has desirable repository characteristics (see FAQ).
- If the repository charges a fee for storing your data, it will typically have a one-time data publishing cost that can be paid during the NIH project period and allow for sharing beyond the project period.
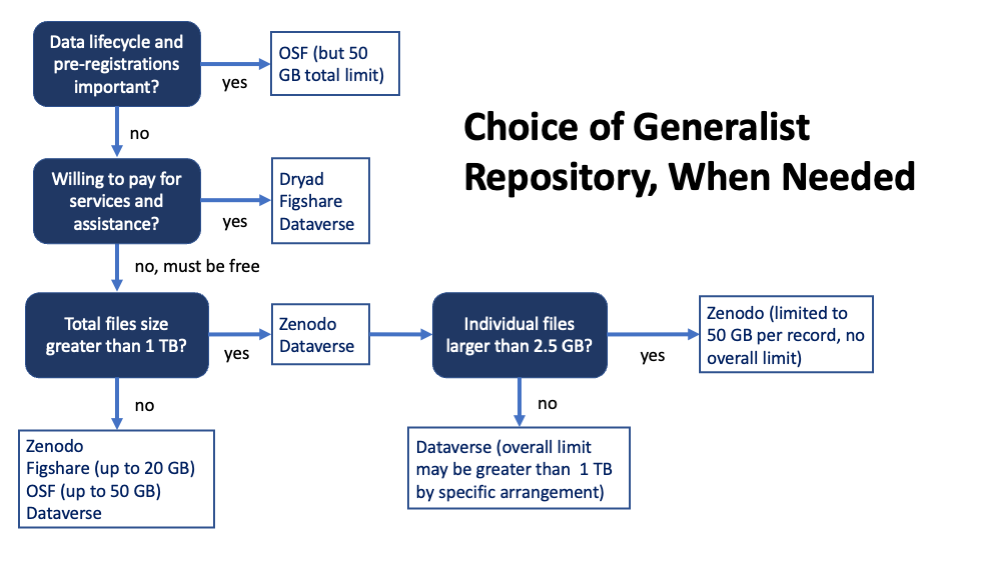
Selecting a Generalist Repository
We have identified 5 generalist repositories that meet the basic requirements of the NIH and are suitable to recommend: Harvard Dataverse, Figshare, Open Science Framework (OSF)*, Dryad, and Zenodo. The three in bold are recommended based on ease of use. OSF is distinct in that it is both easy to use and provides for planning throughout the data life cycle. The Vanderbilt Library provides additional information comparing repositories, and Zenodo published a comparison chart of generalist repositories.
When depositing data into a generalist repository, you will be required to include metadata and, typically, an associated README.txt file to explain the data in the directory. You will also want to cite your data in publications and as a research product in your CV or biosketch. See the FAQs for recommendations on developing each of those items.
FAQs
Special thanks to the Research Integrity & Compliance Subcommittee on NIH Data Management and Sharing Plans; Chuck Sanders, Vice Dean, School of Medicine Basic Sciences; Selene Colon, Assistant Dean of Research Logistics and Compliance, School of Medicine Basic Sciences; Jon Shaw, University Librarian; and Steven Baskauf, Data Science and Data Curation Specialist, Jean and Alexander Heard Libraries.
Last updated 7/1/2024