Project 5 - Systems Engineering and Analysis
Professor John Wikswo is the Co-PI for Project 5 "Systems Engineering & Analysis for Organotypic Culture Models" and director of the Vanderbilt Institute for Integrative Biosystems Research and Education (VIIBRE). Project 5 supports the physiologically realistic function of the mammary, limb, fetal membrane, and liver organotypic culture models (OCMs), operating separately and in combination by coordinating the refinement of SBS-format Integrated Organ Microfluidics (IOM) modules that will provide a common platform architecture for each of the four VPROMPT OCMs. They plan to accomplish this by designing hardware modules that optimize commonality of design between different OCMs, developing readouts to quantify OCM response to toxins and elucidate Adverse Outcome Pathways (AOPs), determining the optimal means to couple a liver OCM to other OCMs, and identifying and circumventing bottlenecks and economic barriers to medium throughput screening (MTS) with OCMs.
Co-Investigators of this effort are Prof. David Cliffel (electrochemical sensors), Prof. John McLean (mass spectrometry), and Prof. Matt Shotwell (biostatistics) at Vanderbilt University.
Background: Depending on the development stage of the OCM, we will provide standalone microfluidic Rotary Planar Peristaltic Micropumps (RPPMs) and multi-port Rotary Planar Valves (RPVs), integrated Perfusion Controller (PC), Microformulator (µF), compact MicroClinical Analyzer (µCA), and custom automation software enabling the scheduling of fluid transfer, media changes, and sample analysis. The µCA electrochemical sensors monitor online measurements of the metabolic state of perfused cultures through detection of key analytes for cellular energy: glucose, lactate, oxygen, and pH. Its programmable pump and valve allow us to perform automated calibrations and sampling.
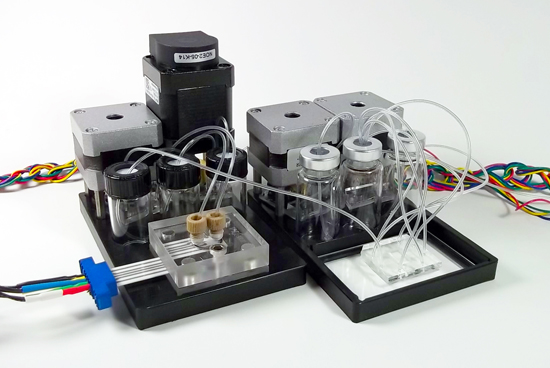

MicroClinical Analyzer and Perfusion Controller Modules
To adequately characterize the complex cellular responses of OCMs to toxins, comprehensive biomolecular profiles must be acquired in a dynamic manner. We have developed on-line methods of sampling cellular secretions from OCMs using ion mobility-mass spectrometry (IM-MS)-based detection.4,5 IM-MS separates biomolecular classes in conformational space based upon respective gas-phase packing efficiencies. This provides a means to perform multiple broad range analyses, or omics experiments, for a given output. Additionally, utilizing accurate mass measurements (<5ppm) and fragmentation spectra, confident molecular identifications can be assigned to features of interest.
We are also sensitive to the issues of targeted versus untargeted searches. The advantage of
the Waters IM-MS/MSE system is that normal IM-MS of intact molecules is interleaved with IM-MS/MSE fragmentation analysis. Typically, the IM separation, the MS mass accuracy, and the use of statistical techniques such as self-organizing heat maps provide a rapid identification of which molecular species in an untargeted search are of statistically significant interest. The MSE data can then be examined without having to rerun the samples and compared to fragmentation atlases to aid in determining the specific molecule that was identified.
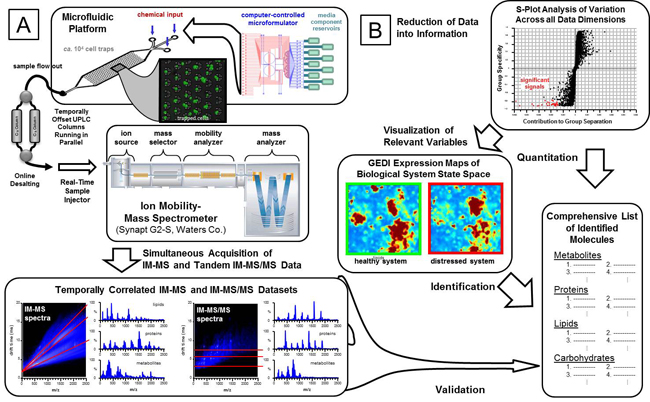
IM-MS workflow overview from microfluidic chip (A) to temporally correlated datasets
| Aim 1 | Design OCM bioreactors that optimize commonality of design. |
| Aim 2 | Develop information-rich readouts to quantify OCM responses to chemical exposure and elucidate Adverse Outcome Pathways (AOPs). |
| Aim 2a | Apply and refine electrochemical sensors of metabolic activity that are matched to the OCMs and culture media and can be applied to MTS. |
| Aim 2b | Mass Spectrometry-Based Analytics. |
| Aim 2c | Refine statistical methodologies for predicting and clustering toxicant responses in OCMs and animal models. |
| Aim 3 | Develop a standardized approach to coupling a liver OCM to other OCMs. |
| Aim 4 | Identify and circumvent bottlenecks and economic barriers to medium throughput screening (MTS). |
