DNA Mechanics
Genome maintenance systems keep cells on track
December 8, 2010 | Leigh MacMillan

Picture a railroad track, running across the countryside. DNA looks something like that track. Its “rails” are chains of chemicals called nucleotides, each pairing with a partner on the opposite chain to form the “ties.” To ensure timely train traffic – and prevent catastrophic derailments – the railroad track needs regular maintenance. So does DNA. Derailments along the DNA track can cause the genetic mutations that lead to cancer.
Our cells employ a range of maintenance “crews” to keep the genome in tip-top shape.
“The fundamental premise of genome maintenance is that DNA is king,” says William Tansey, Ph.D. “Everything that goes on in a cell, absolutely everything, is ultimately programmed as a result of the DNA. The genome has to be maintained in pristine condition. When it’s not, cells acquire mutations and are on their way to becoming cancers.”
Tansey and David Cortez, Ph.D., lead the Genome Maintenance Research Program of the Vanderbilt-Ingram Cancer Center.
The program and its 26 faculty members are seeking to understand “all of the things that have to happen for a cell to make sure that its genome stays the same – within a single cell and after cell division,” says Cortez, Ingram Professor of Cancer Research and professor of Biochemistry.
“All of the things” include the cellular pathways that metabolize carcinogens. They include the teams of proteins that sense and repair DNA damage. They include the processes of DNA replication and cell division. They include the faithful expression of genes. And so on.
“It’s a lot of territory,” admits Tansey, Ingram Professor of Cancer Research and professor and interim chair of Cell and Developmental Biology.
It’s territory that addresses fundamental questions of cell biology – how genetic information is preserved, expressed and passed on.
“Our research program is primarily a basic science program,” Cortez says. “The investigators in the program are interested in the nuts and bolts of these pathways, and how they’re working.”
Because the proteins that keep the genome in fine fettle are highly evolutionarily conserved, the program’s members can take advantage of simpler model organisms – such as bacteria, yeast and fruit flies – for their studies. And their findings, in addition to revealing the workings of the “amazing machines” that cells employ to maintain their genomes, are “critical for clinical cancer care,” Cortez says.
Radiation therapy and many chemotherapy drugs kill cells because they cause DNA damage. The genome maintenance activities in the cancer cells determine if the therapies are effective, or not. Cancer cells can evade DNA-damaging therapies, for example, by having more efficient DNA repair mechanisms or by blocking the signaling pathways that cause cell death in response to DNA damage.
“The more we understand about how genome maintenance pathways are working – in normal cells and in cancer cells – the more we’ll understand why cancer treatments work or don’t work,” Cortez says.
“Our challenge is to exploit genome maintenance as a way of curing cancer.”
Broken track
DNA is under constant assault. Chemicals in the environment, chemical products from normal cellular metabolism, ultraviolet radiation from the sun, X-rays, and a number of other insults damage DNA.

William Tansey, Ph.D.
They add chemical “bits” to the track; they kink the ties; they break one or both of the rails.
As many as 1 million sites in a human cell’s DNA may be damaged every day. Left unrepaired, these lesions can “derail” the protein machines that roll along the DNA to copy it for protein production or cell division, and lead to mutations – changes in the DNA code. Mutations accumulate over time as cells age, and may eventually allow a cell to escape normal growth controls, ramp up its rate of cell division, and develop into cancer.
DNA damage response teams continuously inspect the genome for damage and initiate repairs. They can halt protein traffic along the DNA rails while repairs are made, and in cases of significant damage, they can push the cellular self-destruct button.
These maintenance crews respond to distinct types of damage: some protein teams look exclusively for a certain chemical bit (each team responding to a different type), while other protein teams respond to DNA breaks.
We’re still learning which proteins respond to DNA damage, and how they work. It’s clear though that these pathways are critical: inherited defects often increase a person’s risk for developing cancer.
“Almost all cancers have at their root a genome maintenance problem,” Cortez says. “In order to become a cancer, a cell has to have not one, not two, but several genetic abnormalities, and it needs several more to become aggressive and deadly. That really only happens when one of these systems that prevent mutations is inactivated.”

David Cortez, Ph.D.
Consider BRCA1 and BRCA2, for example. These two “breast cancer susceptibility” genes were discovered in a search for genes associated with inherited breast cancer. A woman who inherits a harmful mutation in either of these genes is about five times more likely to develop breast cancer than a woman who does not have such a mutation. And she may also be at increased risk for ovarian cancer. Men who inherit BRCA1 or BRCA2 mutations are also at increased risk for breast and possibly other types of cancer.
At the time of their discovery, it was unclear how mutations in the BRCA1 and BRCA2 genes contribute to breast cancer development. We know now that the BRCA1 and BRCA2 proteins are DNA repair proteins – they repair breaks that occur in both rails of the DNA chain (double strand breaks), and they coordinate the signaling response to this type of DNA damage.
“So the reason women with inherited mutations in these genes get breast cancer is because they have a decreased ability to maintain the genome, so they’re prone to developing more mutations,” Cortez explains.
The same is true for hereditary colon cancer and for other inherited predispositions for cancer: a person’s cells are “genetically unstable” because of a mutation in a genome maintenance pathway that then generates more mutations.
There is increasing interest in taking advantage of maintenance defects like BRCA1 mutations to selectively target and kill cancer cells with these defects, Cortez says. And recent studies suggest the strategy may be a good one.
Doubling the damage
A protein enzyme called PARP1 – and its signaling pathway – identifies and repairs single strand DNA breaks caused by chemotherapy drugs, ultraviolet light and products of cellular metabolism.
Left unrepaired, single strand breaks can become double strand breaks, which are particularly problematic for cells as they can result in the breaking of chromosomes (and the chromosomal abnormalities that cause cancers and other diseases).
Investigators reasoned that blocking the PARP pathway might “sensitize” cells to the DNA-damaging effects of chemotherapy or radiation – if chemotherapy or radiation alone did not cause enough DNA damage to make the cells self-destruct, then blocking the PARP pathway might help “push cancer cells over the edge,” Cortez says.
And similarly, since cancer cells with a deficiency in BRCA1 or BRCA2 are already inefficient in repairing double strand breaks, then perhaps blocking an additional DNA repair pathway – like PARP – would kill those cells even without DNA-damaging therapies.
PARP inhibitors killed cultured cancer cells and cancers in animal models, and they are now being studied in clinical trials.
Part of a Phase I trial of the PARP inhibitor olaparib in patients with solid tumors evaluated anti-tumor activity in patients with hereditary BRCA-deficient tumors (breast, ovarian or prostate). The tumors shrank or were stabilized in 12 of 19 of these patients.
Olaparib and other PARP inhibitors are now being evaluated in Phase II trials as single agents for BRCA-deficient tumors and in combination with DNA-damaging chemotherapies such as cisplatin and gemcitabine for a variety of tumors.
The effectiveness of PARP inhibitors in killing BRCA-deficient tumors validates a concept called “synthetic lethality.” The idea, Cortez explains, is that a defect in one of two DNA repair pathways (PARP inhibition or BRCA mutation) is not lethal to the tumor cells on its own. But a defect in both pathways at the same time kills the cells.
PARP inhibitors have low toxicity for normal cells, even in patients with an inherited BRCA mutation, because the normal cells in these patients still have one non-mutated BRCA gene that can produce functional repair protein.
“The ongoing development of PARP inhibitors demonstrates how we’re using our knowledge of genome maintenance pathways to specifically target certain kinds of cancer cells – in this case, those with a BRCA deficiency,” Cortez says. “It’s a great example of personalized cancer medicine where we can give treatments with lower toxicities to patients based on the particular changes in their tumors.”
Cortez and his colleagues are conducting studies to discover other “synthetic lethal” relationships like the one between PARP and BRCA that could be exploited therapeutically.
In one series of studies, the investigators treated cultured cells with a PARP inhibitor and used a gene-silencing technology to “turn off” one gene at a time (they tested a series of 73 candidate genes). If they turned off a gene in a synthetic lethal relationship with PARP, they reasoned, the cells would die. They discovered that silencing a gene called USP11 killed the PARP inhibitor-treated cells.
“We didn’t know anything about USP11, but we knew from the screen that it had this relationship with PARP. So we thought maybe it works – like BRCA – in double strand break repair. And it turns out it does,” Cortez says. “So you can also use this synthetic lethal relationship to discover new genes in the pathways.”
Chromatin control
In our cells, DNA does not stretch out long and smooth like railroad tracks. Instead, it condenses itself around histone proteins into a tightly wrapped and wound form, called chromatin.
“If you stretched out all the DNA in a single cell and laid it end to end, it would be about 3 feet long,” says Scott Hiebert, Ph.D., Ingram Professor of Cancer Research and professor of Biochemistry. “And that DNA has to be packaged into the very small space of the nucleus. It’s a really fascinating physical problem for the cell to compact all of its DNA into the nucleus, and then to be able to replicate it and copy parts of it for protein production.”
The cell makes DNA available for copying, at least in part, by modifying histone proteins – adding or removing chemical bits. The addition of chemical “acetyl” groups to histones opens sections of the DNA, making it available to replication, transcription (for protein production), and repair machinery. Removal of the acetyl groups, by proteins called HDACs (histone deacetylases) returns the DNA to a compacted – closed for business – state.
Chemicals that block HDACs are now emerging as a new class of anti-cancer compounds. The HDAC inhibitors vorinostat (Zolinza) and romidepsin (Istodax) are recently FDA-approved treatments for cutaneous T cell lymphomas. More than 10 other HDAC inhibitors are being tested in clinical trials as treatments for a range of cancer types.
HDAC inhibitors kill cancer cells by altering gene expression patterns, causing DNA damage, and interfering with DNA repair. And they may cause widespread disruption of chromatin structure, recent studies by Hiebert and colleagues suggest.
To explore the roles of individual HDACs (there are at least 17 such proteins), Hiebert’s team has used genetic strategies in mice to eliminate the HDAC3 gene.
They found previously that mice without HDAC3 die before birth. In recent studies, they eliminated HDAC3 specifically in the liver and discovered that the mice develop liver cancer.
In cells missing HDAC3, the DNA was “completely open,” Hiebert says. The chromatin no longer existed in its compact/closed form. “Eliminating HDAC3 had a global effect on chromatin, which was completely unexpected.
“We think that removal of HDAC3 deregulates chromatin structure, which then causes DNA damage, genomic instability, and ultimately, cancer. It looks like the target of HDAC inhibitors may really be chromatin structure, but we’re still at a very early stage of sorting out how these drugs are working.”
Although eliminating HDAC3 caused cancer in the mice, Hiebert says that he doesn’t expect HDAC inhibitors will have this adverse effect, because they only block HDAC for short periods of time during therapy.
Full steam ahead
Dineo Khabele, M.D., a gynecologic oncologist, hopes HDAC inhibitors will offer a way to improve treatments for ovarian cancer.

Scott Hiebert, Ph.D., and Dineo Khabele, M.D., are working together to understand the mechanisms of DNA packaging and how those processes can be targeted to improve cancer treatments.
Ovarian cancer is usually diagnosed late, when the disease has already spread outside the ovaries. Although some progress has been made in extending the lives of patients with ovarian cancer, “the overall prognosis is still very poor,” says Khabele, assistant professor of Obstetrics and Gynecology, and Cancer Biology. “We really need to make a bigger impact in overall survival.”
Khabele says being on the “front lines” treating ovarian cancer and seeing the impact of current treatments has motivated her to find something better.
That’s where HDAC inhibitors may come in.
One of the main treatments for ovarian cancer is the chemotherapy drug cisplatin, which kills cells by damaging DNA. Ovarian cancers as a whole may be more likely to have defects in DNA repair pathways that make them more sensitive to DNA-damaging treatments like cisplatin, Khabele says. About 10 percent of ovarian cancers are caused by inherited BRCA1 and BRCA2 mutations, and a greater number appear to have a “BRCA-like” profile of defective DNA repair.
Khabele demonstrated in previous studies that HDAC inhibitors alone or in combination with cisplatin are very effective at killing cultured ovarian cancer cells and ovarian cancer in mouse models.
She and her colleagues are now conducting a retrospective study to search for a “DNA repair defect” profile in preserved tumor tissue from ovarian cancer patients treated at Vanderbilt. And they’re developing new mouse models of ovarian cancer from tumor samples of patients in treatment. They will test HDAC inhibitors in combination with cisplatin to treat the cancers in these mice.
“We’re looking for clues in the tumor profiles and in responses in the mice about which tumors respond best to platinum therapies in combination with HDAC inhibitors,” Khabele says. The investigators believe those will be the tumors with defects in DNA repair.
The ultimate goal, she says, is to use these clues to deliver personalized treatments to ovarian cancer patients – and to have those treatments extend overall survival.
“We’re still in early stages,” Khabele says. “The basic science work to really understand these genome maintenance pathways is critical; it’s absolutely necessary for us to be able to take these ideas to the next level and figure out how to design better drugs, better treatments.”
Vanderbilt-Ingram’s Genome Maintenance Research Program is perfectly situated to meet this challenge, says co-director Tansey.
“We have this island of very basic genome maintenance research going on in a sea of translational science,” he says. “If there ever was an environment for making the leap from the basic findings to clinical care, it’s within the Cancer Center here.”
***
Genome gandy dancers
Our knowledge of genome maintenance systems has exploded in the last 10 years. But we don’t yet know the identity of all of the relevant players, says David Cortez, Ph.D., co-director of the Genome Maintenance Research Program at the Vanderbilt-Ingram Cancer Center.
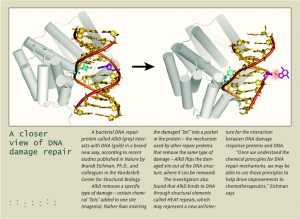
Cortez and colleagues are looking for new genome gandy dancers, proteins that – like the teams of manual laborers who maintained railroad tracks in the past – move through a carefully choreographed routine to maintain DNA.
Their strategy for finding these proteins, Cortez says, is beautifully simple. Using antibodies that bind to sites of DNA damage and that are “hooked to a molecular flashlight,” the investigators can “see” DNA damage in cells.
“The fact that you can visualize DNA damage just by looking through a microscope is so cool. You can’t see the DNA, but you can see these spots light up and know that’s where the damage is,” Cortez says.
The investigators set up a genetic screen in cells to study genes one by one. They used a gene-silencing tool called RNAi to inactivate genes and asked: is there an increase in the amount of DNA damage in the cell? A yes response suggests that the inactivated gene normally plays a role in genome maintenance.
Graduate student Courtney Lovejoy screened 6,386 different RNAi molecules that targeted 2,287 different genes (the genes were selected because they encoded proteins with structural domains associated with nuclear regulatory activities).
The team identified about 100 genes whose inactivation caused increased DNA damage. About half of these were known genes, but the others have not been characterized.
“I’m excited – we have all of these new genes to study and figure out how they’re involved in DNA repair,” Cortez says.
Photo(s) By: Illustration by Brad Yeo | Photos by Joe Howell and John Russell